Category Archives: Programming
Importing GloVe Embeddings into Tensorflow
Machine Learning Crash Course from Google
Earlier this month Google made their internal Machine Learning Crash Course available. You can read more about it on their developer blog. I have a few machine learning projects going, mostly to learn but also to create an alignment-free sequence origin-identification tool. The (unorganized, incomplete) code is available at my GitHub repository. I’m curious about methods to improve genome… Read More »
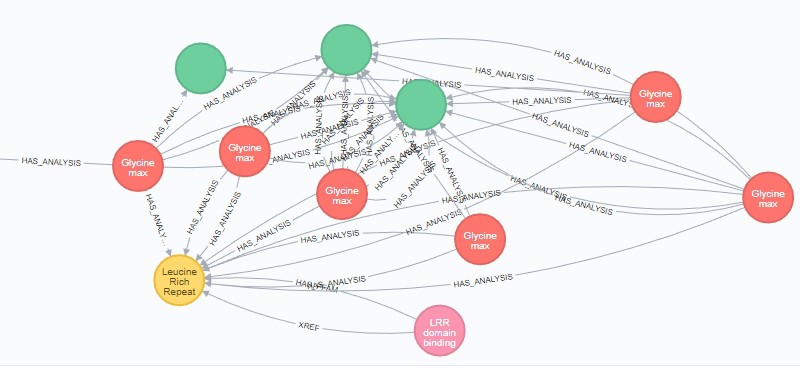
Using ODG from the Neo4j Web Console
The ODG query interface should suffice for many operations, and the command-line interface supports only certain analyses. If you have more advanced queries to run, you can interact with ODG’s generated database from nearly any programming language, using a library or package, via the REST API, or through Neo4j’s Web Console. This tutorial will cover accessing it via… Read More »
Bio* Library for Clojure.
Biotools is my basic bioinformatics file parsing library. You can find it at GitHub. It can parse BLAST+ Tab output (-outfmt “6 std qlen slen”), ExPASY ENZYME.dat, FASTA, GFF3, FPKM Tracking files from Cufflinks, Interproscan tab delimited output, Gene Ontology/Plant Ontology OBO format (any Ontology in OBO format), PMN Pathways format, and a PSI-MITAB 2.5 format. These are all… Read More »
Reading genes.fpkm_tracking into Clojure/Incanter
I needed to analyze a large batch of samples (~300) of genes.fpkm_tracking files in Clojure and Incanter. This guide will show you how I read the files in, only looked at the FPKMs, and converted it into a single dataset. You need a project.clj file somewhere with the dependencies below (incanter, me.raynes.fs).
Experimenting with Pulsar in Clojure
I’ve now started using Pulsar instead of trying out core.async as I needed a lightweight barrier to entry. The problem: My database of choice (Neo4j) takes batch insertions in a single thread only, but clojure is by its very nature multi-threaded/concurrent/parallel (the exact wording of which I am no longer certain!). I process many files when building the… Read More »
Graph Database example using Gene Ontology – Part 1
The Gene Ontology project is a useful tool for anyone doing genomics. It’s a highly relational and controlled vocabulary, making it ideal for use in a graph database. In this example I will show you what a graph database is, and throughout this series we will create a graph database of GO terms, properly linked, inside the Neo4j… Read More »